Chapter 17 – Terminologies (2nd Ed.)
(This Chapter comes from the 2003 2nd Edition of the book and in the 2015 3rd edition has been updated to reflect the many changes to important terminology systems in the last decade)
Chapter 17 – Healthcare terminologies and classification systems
The terms disease and remedy were formerly understood and therefore defined quite differently to what they are now; so, likewise, are the meanings and definitions of inflammation, pneumonia, typhus, gout, lithiasis, &c., different from those which were attached to them thirty years ago…It is evident … that great mischief will in most cases ensue if, in such attempts at definition and explanation, greater importance is attached to a clear and determinate, than to a complete and comprehensive understanding of the objects and questions before us. In a field like ours, clearness can in general be purchased only at the expense of completeness and therefore truth.
Oesterlen, Medical Logic, (1855)
Coding and classification systems have a long history in medicine. Current systems can trace their origins back to epidemiological lists of the causes of death from the early part of the eighteenth century. François Bossier de Lacroix (1706-1777) is commonly credited with the first attempt to classify diseases systematically (ICD-10, 1993). Better known as Sauvages, he published the work under the title Nosologia Methodica.
Linnaeus (1707-1778) who was a contemporary of Sauvages also published his Genera Morborum in that period. By the beginning of the nineteenth century, the Synopsis Nosologiae Methodicae, published in 1785 by William Cullen of Edinburgh (1710-1790) was the classification in most common use.
It was John Graunt who, working about a hundred years earlier, is credited with the first practical attempts to classify disease for statistical purposes. Working on his London Bills of Mortality, he was able to estimate the proportion of deaths in different age groups. For example, he estimated a 36% mortality for liveborn children before the age of 6. He did this by taking all the deaths classified as convulsions, rickets, teeth and worms, thrush, abortives, chrysomes, infants, and livergrown. To these he added half of the deaths classed as smallpox, swinepox, measles, and worms without convulsions. By all accounts his estimate was a good one (ICD-10, 1993).
It has only been in the last few decades that these terminological systems have started to attract wide-spread attention and resources. The ever growing need to amass and analyse clinical data, no longer just for epidemiological purposes, has provided considerable incentive and resources for their development. Further, with the development of computer technology, there has been a belief that such wide-spread collection and analysis of data are now possible. In parallel, the requirement for clinicians to participate in that data collection has meant that they have had more opportunity to work with terminologies, and begin to understand their benefits and limitations.
In the previous chapter, the basic concepts of term, code, and classification were introduced. In this chapter, several of the major coding and classification systems in routine use in healthcare will be introduced, and their features compared. Some specific limitations of each system will be highlighted. In reality there are a large number of such systems in development and use, and they cannot all be identified here. The systems discussed are however representative of most systems in common use, and can serve as an introduction to them. Throughout, a historical perspective will be retained, since in this case the lessons of the past have deep implications for the present. The more general limitations of all terminological systems will be addressed in the following chapter.
17.1 The International Classification of Diseases
Purpose. The International Classification of Diseases (ICD) is published by the World Health Organisation (WHO). Currently in its tenth revision (ICD-10), its goal is to allow morbidity and mortality data from different countries around the world to be systematically collected and statistically analysed. It is not intended, nor is it suitable, for indexing distinct clinical entities (Gersenovic, 1995). The International Nomenclature of Diseases (IND) provides the set of recommended terms and synonyms that correspond to the entries classified in the ICD codes.
History. The ICD can trace its ancestry to the early days of healthcare terminologies. William Farr (1807-1883) became the first medical statistician for the General Register Office of England and Wales. Upon taking office, he found the Cullen classification in use, but that it had not been updated in accordance with medical advances, nor did it seem suitable for statistical purposes. In his first Annual Report of the Registrar General, he noted:
‘The advantages of a uniform statistical nomenclature, however imperfect, are so obvious, that it is surprising that no attention has been paid to its enforcement in Bills of Mortality. Each disease has, in many instances, been denoted by three or four terms, and each term has been applied to as many different diseases: vague, inconvenient names have been employed, or complications have been registered instead of primary diseases. The nomenclature is of as much importance in this department of enquiry as weights and measures in the physical sciences, and should be settled without delay. (ICD-10, 1993).’
Farr toiled hard at improving the classification, and by 1855, the International Statistical Congress adopted a classification based on the work of Farr, and Marc d’Espine of Geneva. Subsequently steered by Jaques Bertillon, this developed into the International List of Causes of Death. This was adopted in 1893, and continued to develop through the turn of the century and beyond, and ultimately evolved into the current ICD system.
In particular, the system was expanded to include not just causes of death, but diseases resulting in measurable morbidity. This expansion started with the urging of Farr. It was supported by Florence Nightingale, who in 1860 urged the adoption of Farr’s disease classification for the tabulation of hospital morbidity in her paper Proposals for a uniform plan of hospital statistics. In 1900 at the First International Conference to revise the Bertillon Classification, a parallel classification of diseases for use in statistics of sickness was finally adopted.
Level of acceptance and use. The ICD today is used internationally by WHO for comparison of statistical returns. It is also adopted by many individual countries in the preparation of their statistical returns. Most other major classification systems endeavour to make their systems compatible with ICD, so that data coded in these systems can be mapped directly to ICD codes. ICD thus acts as a defacto reference point for many healthcare terminologies.
Classification structure. The ICD-10 is a multiple-axis classification system. At its core, the basic ICD is a single list of three alphanumeric character codes. These are organised by category, from A00 to Z99 (excluding U codes which are reserved for research, and for the provisional assignment of new diseases of uncertain aetiology). This level of detail is the mandatory level for reporting to the WHO mortality database and for general international comparisons.
The classification is structured into 21 chapters, and the first character of the ICD code is a letter associated with a particular chapter (Table 17.1).
| Table 17.1: The ICD-10 chapter headings (adapted from ICD-10, 1993). |
| Chapter I | Infectious and parasitic diseases |
| Chapter II | Neoplasms |
| Chapter III | Diseases of the blood and blood forming organs and certain disorders affecting the immune mechanism |
| Chapter IV | Endocrine, nutritional and metabolic diseases |
| Chapter V | Mental and behavioural disorders |
| Chapter VI | Diseases of the nervous system |
| Chapter VII | Diseases of the eye and adnexa |
| Chapter VIII | Diseases of the ear and mastoid process |
| Chapter IX | Diseases of the circulatory system |
| Chapter X | Diseases of the respiratory system |
| Chapter XI | Diseases of the digestive system |
| Chapter XII | Diseases of skin and subcutaneous tissue |
| Chapter XIII | Diseases of musculoskeletal system and connective tissue |
| Chapter XIV | Diseases of the genitourinary system |
| Chapter XV | Pregnancy, childbirth and the puerperium |
| Chapter XVI | Certain conditions originating in the perinatal period |
| Chapter XVII | Congenital malformations, deformations and chromosomal abnormalities |
| Chapter XVIII | Symptoms, signs and abnormal clinical and laboratory findings |
| Chapter XIX | Injuries, poisoning and certain other consequences of external causes |
| Chapter XX | External causes of morbidity and mortality |
| Chapter XXI | Factors affecting health status and contact with health services of a person not currently sick |
Within chapters, the 3 character codes are divided into homogenous blocks reflecting different axes of classification. In Chapter I for example, the blocks signify the axes of mode of transmission and of the broad group of the infecting organism. Within Chapter II on neoplasms, the first axis is the behaviour of the neoplasm, and the next is its site. Within all blocks some codes are reserved for conditions not specified elsewhere in the classification.
When more detail is required, each category in ICD can be further subdivided, using a fourth numeric character after a decimal point, creating up to 10 subcategories. This is used, for example, to classify histological varieties of neoplasms. A few ICD chapters adopt five or more characters to allow further subclassification along different axes.
Since ICD continues to be used for ever-wider applications beyond its intent, the WHO decided in the 10th revision to develop the concept of a family of related classifications surrounding this core set. This ‘family’ contains lists that have been condensed from the full ICD, and lists expanded for speciality-based adaptations (Figure 17.1). It also contains lists that cover topics beyond morbidity and mortality. For example, there are classifications of medical and surgical procedures, disablement and so forth (Gersenovic, 1995).
|
Figure 17.1: The ICD family of disease and health-related classifications (adapted from ICD-10, 1993). |
The International Classification of Functioning, Disability and Health (ICF) is a more recent member of the ICD ‘family’. While ICD-10 focuses on classifying a patient’s diagnosis, ICF is aimed at capturing a description of their capacity to function. ICF describes how people live with their health condition and describes body functions and structures, activities and participation. The domains are classified from body, individual and societal perspectives. Since an individual’s functioning and disability occurs in a context, ICF also includes a list of environmental factors. The ICF is intended to assist with measuring health outcomes.
Limitations. The ICD has developed as a practical, rather than theoretically based, classification. There have been compromises between classification based on axes of aetiology, anatomical site and so on. There have also been adjustments made to it to meet the needs of different statistical applications beyond morbidity and mortality, for example social security. As such, the ICD exists as a practical attempt at compromise between various health care needs. Consequently, for many applications, finer levels of detail may still be needed, or other axes of classification required.
17.2 Diagnosis Related Groups
Purpose. Diagnosis Related Groups (DRGs) relate a patient’s diagnosis and treatment to the cost of their care (Murphy-Muth, 1987; Feinstein, 1988). Developed in the United States by the Health Care Finance Administration, DRGs were designed to support the calculation of federal reimbursement for healthcare delivered through the U.S. Medicare system.
A patient’s principal diagnoses and the procedures they are treated with during hospital admission are used to select the group in the DRG classification that most appropriately describes they overall type of care that has been delivered. Next the group selected is associated with a typical cost. Specifically, DRG funding requires the use of a cost weighting that is applied by the funding agency to determine the actual amount that should be paid to an institution for treating a patient with a particular DRG. The weightings are determined by a formula that is typically developed on a state or national basis.
DRGs are also used to determine an institution’s overall case-mix. The case-mix index helps to take account of the types of patient an individual institution sees, and estimates their severity of illness. Thus a hospital seeing the same proportion of patients as another, but dealing with more severe illness, will have a higher case-mix index. An institution’s case-mix index can then be used in the formula that determines reimbursement per individual DRG. Unsurprisingly different versions of the reimbursement formula favour different types of institution, and case-mix represents an area for ongoing debate and research.
History. In the mid 1970s the Centre for Health Studies at Yale University began work on a system for monitoring hospital utilisation review (Rothwell, 1987). Following a 1976 trial of a DRG system, it was decided to base the final system on the ICD-9-CM which would provide the basic diagnostic categories. The ICD-9-CM (clinical modification) classification was developed from the ICD-9 by the American Commission on Professional and Hospital Activities. It contains finer-grained clinical detail than the old ICD-9, and along with its successors developed in various countries for ICD-10, is intended for healthcare review and reimbursement use.
Level of acceptance and use. DRGs are used routinely in the United States for management review and payment for Medicare and Medicaid patients. Given the importance of reimbursement world-wide, DRGs have undergone ongoing development, and have been adopted in one form or another in many countries outside the USA, including Australia (AR-DRG), Canada (CMG) and countries of Europe and Asia.
Classification structure. Patients are initially assigned a code from ICD-9 CM or a clinical modification of ICD-10. ICD clinical modifications are multiaxial systems closely based on the ICD structure. Diagnoses are then partitioned into one of about 23 Major Diagnostic Categories (MDCs) according to body organ system or disease. The aim of this step is to group codes into similar categories that reflect consumption of resources and treatment (Figure 10.1). The categories are next partitioned based upon the performance of procedures, and on other variables such as the presence of complications and co-morbidities, patient age, and length of stay, before a DRG is finally assigned (Rothwell, 1987). There is thus a process of category reduction at each stage, starting from the many thousands of ICD codes to the few hundred DRGs:
ICD → MDC → DRG
Limitations. Given the local variations in clinical practice, disease incidence, patient selection, procedures performed, and resources, DRGs and case-mix indices will always only give approximate estimates of the true resource utilisation. For example, should a hospital that is developing new and expensive procedures be paid the same amount as an institution that treats the same type of patient with a more common and cheaper procedure? Should quality of care be reflected in a DRG? For example, if a hospital delivers good quality of care that results in better patient outcomes, should it be paid the same as a hospital that performs more poorly for the same type of patient?
As importantly, those institutions that are best able to create DRGs accurately are more likely to receive reimbursement in line with their true expenditure on care. There is thus an implication in the DRG model that an institution actually has the ability to accurately assemble information to derive DRGs and a case-mix index. Given local and national variations in information systems and coding practice, it is likely that institutions with poor information systems will be disadvantaged, unless the information infrastructure across a region is a ‘level playing field’.
Developments. DRGs are designed for use with inpatients. Accordingly, other systems have been developed for other areas of healthcare. Systems such as Ambulatory Visit Groups (AVGs) and Ambulatory Payment Classifications (APCs) have been developed for outpatient or ambulatory care in the primary sector. These are based upon a patient’s diagnosis, intervention, visit status and physician time. Given the increasing age of the population in western nations, there is a tremendous ongoing cost that comes from the chronic care needed by the elderly. Consequently, systems such as Resource Utilisation Groups (RUGs) and the Australian National Sub-Acute and Non-Acute Patient Classification (AN-SNAP) have been developed to help determine the usage of sub-acute and long-term care resources. RUGs are based upon the time spent by nursing home staff when caring for a patient. SNAP includes measures of functional ability.
17.3 The Read codes
Purpose. The Read codes (now simply called the Clinical Terms in the UK) are produced for clinicians, initially in primary care, who wish to audit the process of care. The Clinical Terms Version 3 (CTV3) is intended, like SNOMED International, to code events in the electronic patient record (O’Neil et al., 1995).
History. The Read codes were introduced in the UK in 1986 to generate computer summaries of patient care in primary care. In the subsequent revision Version 2, their structure was changed and based upon ICD-9 and OPCS-4, the Classification of Surgical Operations and Procedures. As Version 2 became increasingly inadequate, the UK’s Conference of Medical Royal Colleges, and the government’s National Health Service (NHS) established a joint Clinical Terms Project, comprising some 40 working groups representing the different specialities. This was subsequently joined by groups representing nurses and allied health professionals. Version 3 of the Read codes was created in response to the output of the Terms project.
Level of acceptance and use. Use of the Read codes is not mandatory in the UK. However, in 1994 it was recommended by the medical and nursing professional bodies as the preferred dictionary for clinical information systems. The Read codes have been purchased by the UK government and made Crown Copyright.
Classification structure. The Read codes have undergone substantive changes through their various revisions, altering not just the classification and terminological content, but also their structure. In Versions 1 and 2, Read was a strictly hierarchical classification system.
Read Version 3 is released in 2 stages and was a ‘superset’ of all previous releases, containing all previous terms, to allow backward compatibility with past versions. Version 3.0 is a kind of compositional classification system. Like SNOMED, a term can appear in several different ‘hierarchical structures’, classified against different axes. Unlike ICD or SNOMED, the codes themselves do not reflect a given hierarchy. They simply act as a unique identifier for a clinical concept. The ‘hierarchy’ exists as a set of links between concepts. Terms can inherit properties across these links. For example, ‘pulmonary tuberculosis’ may naturally inherit from a parent respiratory disorder or a parent infection term.
In Version 3.1, a set of qualifier terms such as anatomical site was added that can be combined with existing terms. When terms are composed, these composites exist outside of any strict hierarchy. To help in the combination of qualifiers with terms, they are grouped into templates. These capture some rules that help describe the range of possible qualifiers that a term in Read can take (Table 17.2).
|
Table 17.2:Example Read Version 3.1 template showing allowable combinations of terms with qualifier attributes, and attribute values (adapted from O’Neil et al., 1995). |
| Object | Applicable Attribute | Applicable values |
| Bone operation | Site | Bone, Part of Bone |
| Fixation of fracture | Reduction method | Percutaneous, open, closed |
| Fixation of fracture using intramedullary nail | Reaming method | Hand, powered rigid, powered flexible, etc. |
| Fixation of fracture using intramedullary nail | Nail Type | Flexible, Locking, Rigid, etc. |
The Read Codes Drug and Appliance Dictionary is part of the Clinical Terms and covers medicinal products, appliances, special foods, reagents and dressings. The dictionary is designed for use in software that requires capture of medication and treatment data such as electronic patient records and prescribing systems.
Like other major systems, Read offers mapping to ICD-9 codes to permit international reporting, and in some cases also provides ICD-10 mapping. A set of Quality Assurance Rules have been developed for the Clinical Terms which are designed to check the clinical, drug and cross-mapping domains between the current and previous versions of the terms and other major terminologies like ICD-10, and for areas of overlap between the domains themselves (Schulz et al., 1998). Each QA rule is written to interrogate the various files that make up the Read Code releases and is designed to identify those concepts or terms that violate the basic structure of the Read Codes.
Although Read Version 3 does not overtly emphasise axes of classification like SNOMED, both systems allow terms to be linked to each other and to inherit properties across those links. Therefore the underlying potential for expressiveness is the same at the structural level. Differences in the number and type of terms, and the richness of interconnections between them are probably greater determinants of difference between these coding systems, than any underlying structural difference. The presence of a fixed hierarchy, as we find with ICD or SNOMED, carries certain benefits of regularity when exploring the system. It also imposes greater constraints when it is necessary to alter the system because of changes to the terminology. In Read, this burden of regularity begins to be shifted to the rules guiding the composition of terms.
Limitations. The Read templates for term composition are limited in their ability to control combination. A much richer language and knowledge base would be needed to regulate term combination (Rector et al., 1995).
17.4 SNOMED
Purpose. The Systematized nomenclature of medicine is intended to be a general-purpose, comprehensive and computer-processable terminology to represent and, according to its creators, will index “virtually all of the events found in the medical record” (Côté et al., 1993).
History. SNOMED was derived from the 1968 edition of the Manual of tumour nomenclature and coding (MONTAC) and the Systematized nomenclature of pathology (SNOP). SNOMED International (or SNOMED III) is a development of the second edition of SNOMED, published in 1979 by the College of American Pathologists (CAP).
Level of acceptance and use. SNOMED is reportedly used in over 40 countries, presumably largely in laboratories for the coding of reports to generate statistics and facilitate data retrieval. Although CAP is a not for profit organisation, in the past SNOMED license fees have often been significant and may have impeded its more widespread adoption.
Classification structure. SNOMED is a hierarchical, multi-axial classification system. Terms are assigned to one of eleven independent systematised modules, corresponding to different axes of classification (Table 17.3). Each term is placed into a hierarchy within one of these modules, and assigned a five or six digit alphanumeric code (Figure 17.2).
|
Table 17.3: The SNOMED International modules (or axes). |
| Module designator |
| Topography (T) |
| Morphology (M) |
| Function (F) |
| Diseases/Diagnoses (D) |
| Procedures (P) |
| Occupations (J) |
| Living Organisms (L) |
| Chemicals, Drugs & Biological Products (C) |
| Physical Agents, Forces & Activities (A) |
| Social Context (S) |
| General Linkage-Modifiers (G) |
Terms can also be cross-referenced across these modules. Each code carries with it a packet of information about the terms it designates, giving some notion of the clinical context of that code (Table 17.4).
|
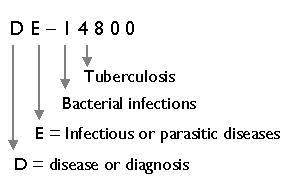
Figure 17.2:SNOMED Codes are hierarchically structured. Implicit in the code, tuberculosis is an infectious bacterial disease. |
SNOMED also allows the composition of complex terms from simpler terms, and is thus partially compositional. SNOMEDInternational incorporates virtually all of the ICD-9-CM terms and codes, allowing reports to be generated in this format if necessary.
|
Table 17.4: An example of SNOMED’s nomenclature and classification. Some terms (e.g. Tuberculosis) can be cross-referenced to others, to give the term a richer clinical context (adapted from Rothwell, 1995). |
| Nomenclature | Classification | ||||||
| Axis | T | + M | + L | + F | = D | ||
| Term | Lung | + Granuloma | + M. tuberculosis | + Fever | = Tuberculosis | ||
| Code | T-28000 | + M-44000 | + L-21801 | + F-03003 | = DE-14800 | ||
SNOMED RT (Reference Terminology) was released in 2000 to support the electronic storage, retrieval and analysis of clinical data (Spackman et al, 1997). A reference terminology provides a common reference point for comparison and aggregation of data about the entire health care process, recorded by multiple different individuals, systems, or institutions. Previous versions of SNOMED expressed terms in a hierarchy that was optimized for human use. In SNOMED RT, the relationships between terms and concepts are contained in a machine-optimised hierarchy table. Each individual concept is expressed using a description logic, which makes explicit the information that was implicit in earlier codes (Table 17.5).
|
Table 17.5:Comparison between implicitly coded information about “postoperative esophagitis” in SNOMED III Codes and the explicit coding in SNOMED RT. (from Spackman et al, 1997) |
| SNOMED III termcode and English nomenclature: | D5-30150 Postoperative esophagitis |
| SNOMED III components of the concept: | T-56000 EsophagusM-40000 InflammationF-06030 Post-operative state |
| Cross-reference field in SNOMED III: | (T-56000)(M-40000)(F-06030) |
| Parent term in the SNOMED III hierarchy: | D5-30100 Esophagitis, NOS |
| Essential characteristics, in SNOMED RT syntax: | D5-30150:D5-30100 &(assoc-topography T-56000) &(assoc-morphology M-40000) &(assoc-etiology F-06030) |
Limitations. It is possible, given the richness of the SNOMED International structure, to express the same concept in many ways. For example, acute appendicitis has a single code D5-46210. However, there are also terms and codes for ‘acute’, ‘acute inflammation’, and ‘in’. Thus this concept could be expressed either as Appendicitis, acute; or Acute inflammation, in, Appendix; and Acute, inflammation NOS, in, Appendix (Rothwell, 1995). This makes it difficult for example, to compare similar concepts that have been indexed in different ways, or to search for a term that exists in different forms within a patient record. The use of description logic in SNOMED RT is designed to solve this problem. Further, while SNOMED permits single terms to be combined to create complex terms, rules for the combination of terms have not been developed. Consequently such compositions may not be clinically valid.
17.5 SNOMED CT (Clinical Terms)
Purpose. SNOMED Clinical Terms is designed for use in software applications like the electronic patient record, decision support systems, and to support the electronic communication of information between different clinical applications. Its designers goal is that SNOMED CT should become the accepted international terminological resource for healthcare, supporting multilingual terminological renderings of common concepts.
| Figure 17.3:Outline of the SMOMED CT core structure (after College of American Pathologists, 2001) |

History. In 1999 the College of American Pathologists and the UK NHS announced their intention to unite SNOMED RT and
Clinical Terms Version 3. The stated intention in creating the common terminology was to decrease duplication of effort and to create a unified international terminology that supports the integrated electronic medical record. SNOMED CT was first released for testing in 2002.
Level of acceptance and use. SNOMED CT supersedes SNOMED RT and the Clinical Terms Version 3. It will gradually replace CTV3 in the UK as the terminology of choice used in the National Health Service (NHS).
Classification structure. The SNOMED CT core structure includes concepts, descriptions (terms) and the relationships between them (Figure 17.3). Like SNOMED-RT and CTV3, SNOMED CT is a compositional and hierarchical terminology. It is multiaxial and utilises description logic to explicitly define the scope of a concept. There are 15 top-level hierarchies (Table 17.6). The hierarchies go down an average of 10 levels per concept.
| Table 17.6: The top-level hierarchies of SMOMED CT. |
| Procedure / intervention includes all purposeful activities performed in the provision of health care. |
| Finding / disorder groups together concepts that result from an assessment or judgment. |
| Measurable / observable entity includes observable functions such as “vision” as well as things that can be measured such as “hemoglobin level”. |
| Social / administrative concept aggregates concepts from the CTV3“administrative statuses” and “administrative values” hierarchies as well as concepts from the SNOMED RT “social context” hierarchy. |
| Body structure includes anatomical concepts as well as abnormal body structures, including the “morphologic abnormality” concepts. |
| Organism includes all organisms, including micro-organisms and infectious agents (including prions), fungi, plants and animals. |
| Substance includes chemicals, drugs, proteins and functional categories of substance as well as structural and state-based categories, such as liquid, solid, gas, etc. |
| Physical object includes natural and man-made objects, including devices and materials. |
| Physical force includes motion, friction, gravity, electricity, magnetism, sound, radiation, thermal forces (heat and cold), humidity, air pressure, and other categories mainly directed at categorizing mechanisms of injury. |
| Event is a category that includes occurrences that result in injury (accidents, falls, etc), and excludes procedures and interventions. |
| Environment / geographic location lists types of environment as well as named locations such as countries, states, and regions. |
| Specimen lists entities that are obtained for examination or analysis, usually from the body of a patient. |
| Context-dependent category distinguishes concepts that have pre-coordinated context, that is, information that fundamentally changes the type of thing it is associated with. For example, “family history of” is context because when it modifies “myocardial infarction”, the resulting “family history of myocardial infarction” is no longer a type of heart disease. Other examples of contextual modifiers include “absence of”, “at risk of” etc. |
| Attribute lists the concepts that are used as defining attributes or qualifying attributes, that is, the middle element of the object-attribute-value triple that describes all SNOMED CT relationships. |
| Qualifier value categorizes the remaining concepts (those that haven’t been listed in the categories above) that are used as the value of the object-attribute-value triples. |
SNOMED CT incorporates SNOMED RT and Clinical Terms Version 3 (Kim and Frosdick, 2001) as well as mappings to classifications such as ICD-9-CM and ICD-10. It is substantially larger than either SNOMED-RT or CTV3, containing over 300,000 concepts, 400,000 terms and more than 1,000,000 semantic relationships. SNOMED CT also integrates LOINC (Logical Observation Identifier Names and Codes) to enhance its coverage of laboratory test nomenclature. Most of the features of the parent terminologies are incorporated into SNOMED CT. For example the CTV3 templates, although not explicitly named in the new structure, are essentially functionally preserved in SNOMED CT.
Limitations: Since SNOMED CT is a compositional terminology, there is strong requirement to prevent illogical compositions being created, and while a form of type checking is implemented, explicit compositional controls are not evident in the early releases of the terminology.
Reviewing a sample of 1,890 descriptions obtained from the initial merging of the two parent terminologies found a 43% redundancy in terms (Sable et al, 2001). While some terms were simply common to both parent systems, many terms were problematic in some way. For example, some terms were either vague or ambiguous, used the logical connectors ‘and’ and ‘or’ incorrectly, had flawed hierarchy links, or contained knowledge about disease processes that should have been beyond the scope of the terminology. Many of these problematic terms were identified automatically, but many others required visual inspection and discussion to be resolved. While the process of merging the two terminologies has substantially improved the quality assurance standard of the resulting terminology, these problems raise many issues fundamental to terminology construction, which are discussed in the following chapter.
17.6 The Unified Medical Language System (UMLS)
Purpose: The UMLS is the Rosetta stone of international terminologies. It links the major international terminologies into a common structure, providing a translation mechanism between them. The UMLS is designed to aid in the development of systems that retrieve and integrate electronic biomedical information from a variety of sources and to permit the linkage of disparate information systems, including electronic patient records, bibliographic databases, and decision support systems. A long-term research goal is to enable computer systems to “understand” medical meaning
History: In 1986, the U. S. National Library of Medicine (NLM) began a long-term research and development project to build a Unified Medical Language System (Humphreys and Lindberg, 1989).
Level of acceptance and use: Broad use of the UMLS is encouraged by distributing it free-of-charge under a license agreement. The UMLS is widely used in clinical applications, and the NLM itself uses the UMLS in significant applications including PubMed and the web-based consumer health information initiative at ClinicalTrials.gov.
Classification structure: The UMLS is composed of three “Knowledge Sources”, a Metathesaurus, a semantic network, and a lexicon (Lindberg et al, 1993).
The UMLS Metathesaurus provides a uniform format for over 100 different biomedical vocabularies and classifications. Systems integrated within the UMLS include ICD-9, ICD-10, the Medical Subject Headings (MeSH), ICPC-93, WHO Adverse Drug Reaction Terminology, SNOMED-II, SNOMED-III, and the UK Clinical Terms. The 2002AD edition of the Metathesaurus includes 873,429 concepts, 2.10 million concept names in its source vocabularies, and over 10 million relationships between them.
The Metathesaurus is organized by concept and does not include an over-arching hierarchy. It can be conceptualised as a web rather than as a hierarchical tree, linking alternative names and views of the same concept together and identifying useful relationships between different concepts. This method of structuring UMLS allows the component terminologies to maintain their original structure within UMLS, as well as linking similar concepts between the component terminologies.
Each concept has attributes that define its meaning, e.g., semantic types or categories to which it belongs, its position in the source terminology hierarchy, and a definition. Major UMLS semantic types include organisms, anatomical structures, biologic function, chemicals, events, physical objects, and concepts or ideas.
A number of relationships between different concepts are represented including those that are derived from the source vocabularies. Where the parent terminology expresses a full hierarchy, this is fully preserved in UMLS. The Metathesaurus also includes information about usage, including the name of databases in which the concept originally appears.
The UMLS is a controlled vocabulary and the UMLS Semantic Network is used to ensure the integrity of meaning between different concepts. It defines the types or categories to which all Metathesaurus concepts can be assigned and the permissible relationships between these types (e.g., “Virus” causes “Disease or Syndrome”). There are over 134 semantic types that can be linked by 54 different possible relationships. The primary link is the `isa’ link, which establishes the hierarchy of types within the Network. A set of non-hierarchical relations between the types includes `physically related to,’ `spatially related to,’ `temporally related to,’ `functionally related to,’ and `conceptually related to.’
The SPECIALIST Lexicon is intended to assist in producing computer applications that need to translate free-form or natural language into coded text. It contains syntactic information for terms and English words, including verbs that do not appear in the Metathesaurus. For example, it is used to generate natural language or lexical variants of words e.g. the word “treat” has three variants that all have the same meaning as far as the Metathesaurus is concerned: treats, treated or treating.
Limitations: The very size and complexity of the UMLS may be barriers to its use, offering a steep learning curve compared to any individual terminology system. Its size also poses great challenges in system maintenance. Every time one of the individual terminologies incorporated into UMLS changes, technically those changes must be reflected in the UMLS. Consequently regular and frequent updates to the UMLS are issued, and as the system grows the likelihood of errors being introduced will increase, as we shall see in the next chapter.
| Table 17.7: A comparison of coding for four different clinical concepts using some of the major coding systems (National Centre for Classification in Health, Australia). |
| Clinical Concept | UMLS | ICD10 | ICD9CM 4th Edition | READ 1999 | SNOMED International 1998 | SNOMED CT 2002 |
| Chronic ischaemicheartdisease | 448589 Chronic ischaemic heart disease | I25.9 Chronic ischaemic heart disease | 414.9 Chronic ischaemic heart disease | XE0WG Chronic ischaemic heart disease NOS | 14020 Chronic ischaemic heart disease | 84537008 Chronic ischaemic heart disease |
| Epidural haematoma | “453700 Hematoma, epidural” | S06.4 Epidural haemorrhage | 432.0 Nontraumatic extradural haemorrhage | Xa0AC Extradural haematoma | 89124 Extradural haemorrhage | 68752002 Nontraumatic extradural haemorrhage |
| Lympho-sarcoma | “1095849 Lymphoma, diffuse” | C85.0 Lymphosarcoma | 200.1 Lymphosarcoma | B601z Lymphosarcoma | “95923 Lymphosarcoma, diffuse” | “1929004 Malignant lymphoma, non-Hodgkin” |
| CommonCold | 1013970 Common cold | J00 Acute nasopharyngitis [common cold] | 460 Acute nasopharyngitis [common cold] | XE0X1 Common cold | 35210 Common cold | 82272006 Common cold |
The richness of the linkages between concepts also offers subtle problems at the heart of terminological science. For example, the‘meaning’ of a UMLS concept comes from its relationships to other concepts, and these relationships come from the original source terminologies. However a precise concept definition from one of the original terminologies like ICD or SNOMED may be blurred by addition of links from another terminology that contains a similar concept (Campbell et al, 1998). For example, “gastrointestinal transit” in the Medical Subject Headings (MeSH) is used to denote both the physiologic function and the diagnostic measure (Spackman et al., 1997).
Since UMLS is not designed to contain an ontology, which could aid with conceptual definition, it is difficult to control for such semantic drift.
17.7 Comparing coding systems is not easy
Unsurprisingly, the same clinical concept might look very different when coded using different classification systems (Table 17.7) The different origins of the systems, and the different revision histories each has had, inevitably result in the use of different terms for similar concepts. While it is beguiling to try to compare the utility of different coding systems, such comparisons are often ill-considered. This is because it is not always obvious how to compare the ability of different systems to code concepts found in a patient record. For example, Campbell et al. (1994), reported results of various systems coding terms found in selected problem lists from US patient records. They assessed that ICD-9-CM and Read Version 2 ‘perform much more poorly for problem coding’ than either SNOMED or the UMLS systems. As a consequence they concluded that ‘both UMLS and SNOMED are more complete than alternative systems’ when developing computer-based patient records.
Such generalisations are not meaningful. Firstly, term requirements vary from task to task. Indeed, terms develop out of the language of particular groups on particular tasks. It is thus not meaningful to compare performance on one task and deduce that similar outcomes will result for tests on other tasks.
As critically, term use will vary between user populations. The terms used in a primary care setting will differ to those used in a clinic allied to a hospital, reflecting different practices and patient populations. Differing disease patterns and practices also distinguish different nations. A system like Read Version 2, designed for UK primary care, may not perform as well in US clinics as a US designed system. The reverse may also be true of a US designed system applied in the UK.
In summary, coding systems should be compared on specified tasks and contexts, and the results should only cautiously be generalised to other tasks and contexts. Equally the poor performance of coding systems on tasks outside the scope of their design should not reflect badly on their intended performance.
Discussion Points
1. How likely is it that a single terminology system will emerge as an international standard for all clinical activities?
2. Take the two terminologies created from the discussion section of the previous chapter, and now merge the two into one common terminology. As you go, note the issues that arise, and the methods you used to settle any differences. Explain the rational (or otherwise) basis for the merger decisions.
3. Are there any clinically significant differences that might arise out of the different codings in Table 17.7? What impact might such differences make on epidemiological surveys of population health?
4. You have been asked to oversee the transition from ICD-9-CM to ICD-10-CM at your institution. What social and technical challenges do you expect to face? How will you plan to deal with them?
5. Many countries will take a major terminology like ICD and customise it to suit their local needs. Discuss the costs and benefits of this approach from an individual country’s point of view. What might the impact of localisation be on the collection of international statistics?
Chapter summary
1. The International Classification of Diseases (ICD) is published by the World Health Organisation. Currently in its tenth revision (ICD-10), its goal is to allow morbidity and mortality data from different countries around the world to be systematically collected and statistically analysed.
2. Diagnosis Related Groups (DRGs) relate patient diagnosis to cost of treatment. Each DRG takes the principle diagnosis or procedure responsible for a patient’s admission, and is given a corresponding cost weighting. This weight is applied according to a formula to determine the amount that should be paid to an institution for a patient with a particular DRG. DRGs are also used to determine an institution’s overall case-mix.
3. The Systematized Nomenclature of Medicine (SNOMED) is intended to be a general-purpose, comprehensive and computer-processable terminology to represent. Derived from the 1968 edition of the Manual of Tumour Nomenclature and Coding, the second edition of SNOMED International is reportedly being translated into twelve separate languages.
4. The Read codes are produced for clinicians, initially in primary care, who wish to audit the process of care. Version 3 is intended, like SNOMED International, to code events in the electronic patient record.
5. Coding systems should be compared on specified tasks, and results should only cautiously be generalised to other tasks, and populations. Equally the poor performance of coding systems on tasks outside their design should not reflect badly on their intended performance.
© Enrico Coiera 1997-2014

Leave a comment